(→additional reading) |
|||
| Line 239: | Line 239: | ||
[http://biomechanics.stanford.edu/me233_20/reading/linka20d.pdf (r15)]<br> | [http://biomechanics.stanford.edu/me233_20/reading/linka20d.pdf (r15)]<br> | ||
| − | linka k et al. is it safe to lift | + | linka k et al. is it safe to lift covid-19 travel bans. the newfoundland story. comp mech doi:10.1007/s00466-020-01899-x. |
[http://biomechanics.stanford.edu/me233_20/reading/linka20c.pdf (r16)]<br> | [http://biomechanics.stanford.edu/me233_20/reading/linka20c.pdf (r16)]<br> | ||
| − | anderson rm, may rm | + | anderson rm, may rm. directly transmitted infectious diseases: control by vaccination, science 215 (1982) 1053-1060 |
[http://biomechanics.stanford.edu/me233_20/reading/anderson82.pdf (r17)]<br> | [http://biomechanics.stanford.edu/me233_20/reading/anderson82.pdf (r17)]<br> | ||
| Line 251: | Line 251: | ||
kuhl e. data-driven modeling of COVID-19 - lessons learned. extr mech lett; doi:10.1016/j.eml.2020.100921. | kuhl e. data-driven modeling of COVID-19 - lessons learned. extr mech lett; doi:10.1016/j.eml.2020.100921. | ||
[http://biomechanics.stanford.edu/me233_20/reading/kuhl20.pdf (r19)]<br> | [http://biomechanics.stanford.edu/me233_20/reading/kuhl20.pdf (r19)]<br> | ||
| + | |||
| + | soper a. the lessons of the pandemic, science xlix (1919) 501-505. [http://biomechanics.stanford.edu/me233_20/reading/liu20.pdf (h01)]<br> | ||
| + | |||
| + | liu y et al., the reproduction number of COVID-19 is higher compared to SARS coronavirus. j travel med (2020) | ||
| + | [http://biomechanics.stanford.edu/me233_20/reading/liu20.pdf (h02)]<br> | ||
| + | |||
| + | oden tj. adaptive multiscale predictive modeling. acta numerica (2018) 353-450. [http://biomechanics.stanford.edu/me233_20/reading/oden18.pdf (h03)]<br> | ||
| + | |||
| + | linka k et al. is it safe to lift covid-19 travel bans. the newfoundland story. comp mech doi:10.1007/s00466-020-01899-x. | ||
| + | [http://biomechanics.stanford.edu/me233_20/reading/linka20c.pdf (p01)]<br> | ||
Revision as of 17:37, 20 August 2020
Contents |
fall 21 - me233 - data-driven modeling of covid-19
|
me 233 - data-driven modeling of covid-19 ellen kuhl
kevin linka fall 2021 |
dissecting brains

| 
| 
| 
|

| 
| 
| 
|

| 
| 
| 
|

| 
| 
| 
|
objectives
understanding the outbreak dynamics of COVID-19 through the lens of mathematical models is an elusive but significant goal. within only half a year, the COVID-19 pandemic has resulted in more than 20 million reported cases across 188 countries with more than 700,000 deaths worldwide. this has generated an unprecedented volume of data; yet, the precise role of mathematical modeling in providing quantitative insight into the COVID-19 pandemic remains a topic of ongoing debate. this course discusses how to design computational tools to understand the COVID-19 outbreak. we focus on mathematical epidemiology, infectious disease models, concepts of the effective reproduction number and herd immunity, network modeling, outbreak dynamics and outbreak control, bayesian methods, model calibration and validation, prediction and uncertainty quantification. we highlight the early success of classical models for infectious diseases and show why these models fail to predict the current outbreak dynamics of COVID-19. we illustrate how data-driven modeling can integrate classical epidemiology modeling and machine learning to infer critical disease parameters—in real time—from reported case data to make informed predictions and guide political decision making. We critically discuss questions that current models can and cannot answer and showcase controversial decisions around the early outbreak dynamics, outbreak control, and exit strategies.
grading
- 15 % homework 01 - mathematical epidemiology of covid-19
- 15 % homework 02 - early outbreak dynamics of covid-19
- 15 % homework 03 - outbreak control of covid-19
- 25 % final project - presentation graded by class
- 30 % final project - report graded by instructors
final project
syllabus
| day | date | topic | slides | read | hw | |
|---|---|---|---|---|---|---|
| tue | sept | 15 | introduction to covid-19 | s01 | ||
| thu | sept | 17 | I. infectious disesaes - a brief history | s02 | r02 | |
| tue | sept | 22 | II. mathematical epidemiology - 1. intro to compartment modeling | s03 | r02 | |
| thu | setp | 24 | II. mathematical epidemiology - 2. compartment models | s04 | r04 | h01 |
| tue | sept | 29 | II. mathematical epidemiology - 3. endemic disease modeling | s05 | r05 | |
| thu | oct | 01 | III. data-driven modeling - 1. compartment modeling of covid-19 | s06 | r06 | |
| tue | oct | 06 | III. data-driven modeling - 2. early outbreak dynamics of covid-19 | s07 | r07 | |
| tue | oct | 08 | III. data-driven modeling - 3. asymptomatic transmission of covid-19 | s08 | r08 | h02 |
| tue | oct | 13 | III. data-driven modeling - 4. inferring outbreak dynamics of covid-19 | s09 | r09 | |
| thu | oct | 15 | IV. modeling outbreak control - 1. managing infectious diseases | s10 | r10 | |
| tue | oct | 20 | IV. modeling outbreak control - 2. change-point modeling of covid-19 | s11 | r11 | |
| thu | oct | 22 | IV. modeling outbreak control - 3. dynamic modeling of covid-19 | s12 | r12 | h03 |
| tue | oct | 27 | V. network modeling - 1. network modeling of epidemiology | s13 | r13 | |
| thu | oct | 29 | V. network modeling - 2. network modeling of covid-19 | s14 | r14 | |
| tue | nov | 03 | V. network modeling - 3. dynamic network modeling of covid-19 | s15 | r15 | |
| thu | nov | 05 | VI. informing political decision making - exit strategies | s16 | r12 | p01 |
| tue | nov | 10 | VI. informing political decision making - vaccination strategies | s17 | r17 | |
| thu | nov | 12 | VI. informing political decision making - the second wave | s18 | r18 | |
| tue | nov | 17 | data-driven modeling of covid-19 - lessons learned | s19 | r19 | |
| thu | nov | 19 | data-driven modeling of covid-19 - final project presentations | s20 |
python files
we'll share the python files for the projects here
homework 01
learn mathematical epidemiology of covid-19
tools solving ordinary differential equations, time integration, python
apps classical SEIR model, alpha, beta, gamma, Ro = beta/gamma
read soper a, the lessons of the pandemic, science xlix (1919) 501-505
- identify three similarities between the pandemic in 1918 and the covid-19 today
- identify three differences between the pandemic in 1918 and the covid-19 today
- discuss, in two sentences for each, why
- implement SEIR model in python
- time integration: plot SEIR over t for alpha = 2.5 /days, gamma = 6.5 /days, beta = 13 /days
- vary the time step size delta t; discuss the results
- sensitivity analysis: plot SEIR over t for alpha = 2.5 /days; gamma = 6.5 /days
- vary beta such that Ro = beta / gamma = [ 2.0, 3.0, 4.0, 5.0 ], discuss the results
homework 02
learn early outbreak dynamics of covid-19
tools data-driven modeling, parameter identification, python
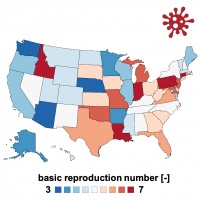
apps basic reproduction number Ro = beta/gamma
read liu y et al., the reproduction number of covid-19. j travel med (2020)
- identify the range of the basic reproduction number for covid-19
- compare it against classical infectious diseases and other coronavirus diseases
- download covid-19 data for your home country, state, or city from an online database
- plot the time evolution of I and R
- fit the static SEIR model from homework 01 to the data
- advise: you may fix alpha and gamma, and only fit beta and eta=N*/N
- plot the recorded I and R and the modeled SEIR over t
- discuss your results, how good is the fit, when is it best and why
- calculate the basic reproduction number Ro = beta / gamma
- compare your Ro against other reported values for covid-19 and other infectious diseases
- for your Ro, calculate the herd immunity threshold
homework 03
learn outbreak control of covid-19
tools combining machine learning and mechanistic modeling, python + PyMC3
apps effective reproduction number R(t) = beta(t) / gamma
read oden tj, adaptive multiscale predictive modeling. acta numerica (2018) 353-450
- explain bayes rule, eq (4.1), in your own words
- interpret its four terms in relation to this homework
- download covid-19 data for your home country, state, or city from an online database
- fit the dynamic SEIR model to the data
- combine dynamic SEIR model with Bayesian methods to learn R(t) for the same data as in homework 02
- plot the recorded I and R and the modeled SEIR over t
- discuss your results, how does the fit differ from homework 02
- plot the effective reproduction number R(t) = beta(t) / gamma
- plot and discuss confidence intervals
- comment on trends, e.g., shelter in place, lockdown, travel restrictions
final project
learn informing decision making
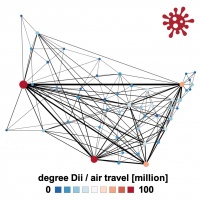
tools solving partial differential equations, network modeling, python + PyMC3
apps dynamic network SEIR model
read linka k et al. is it safe to lift covid-19 travel bans. comp mech. doi:10.1007/s00466-020-01899-x
- guided by the newfoundland example, develop your own model for stanford campus reopening
- create an influx network, e.g., use origin of incoming students, by country or state
- estimate influx dynamics, select an epidemiology model, e.g., SIR or SEIR
- use current dynamics of incoming locations from open source database or model
- estimate number of incoming infectious frosh, sophomores, juniors, seniors
- forecast SEIR populations on campus and discuss confidence intervals
- make informed recommendations of gradual campus reopening
additional reading
bar-on et al., sars-cov-2 (covid-19) by the numbers, elife 9 (2020) e57309.
(r02)
bauer f, compartment models in epidemiology, mathematical epidemiology (2008) 19-79.
(r03)
hethcode hw, the mathematics of infectious disease, siam review 42 (2020) 599-653.
(r04)
dietz k, the estimation of the basic reproduction number for infectious diseases, stat meth med res 2 (1993) 23-41.
(r05)
peirlinck m, et al. outbreak dynamics of covid-19 in china and the united states. medRxiv: doi:10.1101/2020.04.06.20055863.
(r06)
park et al., reconciling early-outbreak estimates of the basic reproduction number and its uncertainty. j royal soc interface 17 (2020) 20200144.
(r07)
ioannis j, the invection fatality rate of covid-19 inferred from seroprevalence data, medRxiv, doi:10.1101/2020.05.13.20101253.
(r08)
peirlinck m et al., visualizing the invisible: the effect of asymptomatic transmission, medRxiv, doi: 10.1101/2020.05.23.20111419
(r09)
wilder-smith a, freedman do. isolation, quarantine, social distancing
and community containment, j travel med (2020) 1-4.
(r10)
dehning et al., inferring change points in the spread of COVID-19 reveals the effectiveness of interventions, science doi:10.1126/science.abb9789.
(r11)
linka et al., the reproduction number of covid-19 and its correlation with public health interventions, comp mech, doi:10.1007/s00466-020-01880-8.
(r12)
pastor-satorras r et al., epidemic processes in complex networks, rev mod phys 87 (2015) 926-973.
(r13)
linka k et al. outbreak dynamics of covid-19 in europe and the effect of travel restrictions. comp meth biomech biomed eng; 2020; 23:710-717.
(r14)
linka k et al. global and local mobility as a barometer for COVID-19 dynamics. medRxiv doi:2020.06.13.20130658.
(r15)
linka k et al. is it safe to lift covid-19 travel bans. the newfoundland story. comp mech doi:10.1007/s00466-020-01899-x.
(r16)
anderson rm, may rm. directly transmitted infectious diseases: control by vaccination, science 215 (1982) 1053-1060
(r17)
grassly nc, fraser c, seasonal infectious disease epidemiology, proc royal
soc b 273 (2006) 2541-2550.
(r18)
kuhl e. data-driven modeling of COVID-19 - lessons learned. extr mech lett; doi:10.1016/j.eml.2020.100921.
(r19)
soper a. the lessons of the pandemic, science xlix (1919) 501-505. (h01)
liu y et al., the reproduction number of COVID-19 is higher compared to SARS coronavirus. j travel med (2020)
(h02)
oden tj. adaptive multiscale predictive modeling. acta numerica (2018) 353-450. (h03)
linka k et al. is it safe to lift covid-19 travel bans. the newfoundland story. comp mech doi:10.1007/s00466-020-01899-x.
(p01)